Nucleotides form nucleic acids
ATP
DNA and RNA
DNA replication
The genetic code
Protein synthesis
Nucleotides form nucleic acids
Nucleic acids are polymers of the nucleotide monomer and include the central biological molecules DNA (deoxyribonucleic acid), RNA (ribonucleic acid) and ATP (adenosine triphosphate), playing roles in inheritance, protein synthesis and metabolism.
ATP is adenosine triphosphate, a nucleotide involved in metabolism and many biological processes (the molecular energy currency).
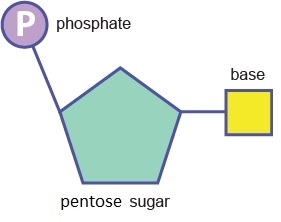
A nucleotide is a molecule composed of a pentose sugar, an organic base and at least one phosphate group.
Chemical energy is important in biological processes because it helps maintain the many reactions that go against a natural equilibrium. These include movement of chemicals against a concentration gradient via active transport, enabling enzyme action as ATP acts as a coenzyme, and in a certain cells, movement.
As I write this I am absolutely knackered which is juuuuuuuuust hilarious as I am about to cover ATP! Adenosine triphosphate is a small molecule whose constant breaking down and putting back together reactions form the basis of our biological processes which require chemical energy.
ATP
As for many of the different other chemicals that we have covered such as carbohydrates and nucleic acids, these ATP reactions are condensation and hydrolysis. However, here we are not talking about monomers forming polymers or polymers breaking back into monomers. We are talking about adenosine triphosphate breaking down into..
