Amino acids form peptide bonds
Identifying amino acids through chromatography
Protein structure
Enzyme structure and function
Proteins in blood clotting, medicine and blood donation
Amino acids form peptide bonds
Proteins are at the heart of living organisms. Their functions are very varied, from the hair on your head, to the haemoglobin in your red blood cells (which carries oxygen around the body), to the claws of a lion, to insulin (blood glucose regulation). All these highly varied proteins are made of their building blocks – amino acids. This is what the generalised structure of an amino acid looks like (make sure you can draw this):
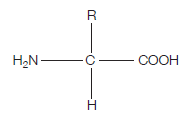
If you’re wondering what this actually is, read on. The clues are in the name (as they usually are).
AMINO – the H2N on the left hand side is an amino group.
ACID – the COOH on the right hand side is a carboxylic acid group (simply an acid) The hydrogen (H) on the bottom is there all the time (just like the amino group and the acid group), while the R group is the variable which determines what particular amino acid this will be.
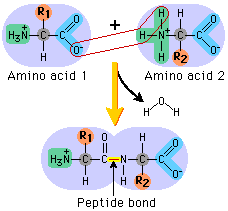
For example, if the R group was a hydrogen, the amino acid would be glycine. The next diagram shows condensation, and the subsequent formation of a bond between two amino acids (any two). This bond is a peptide bond. The resulting molecule is called a polypeptide.
Identifying amino acids through chromatography
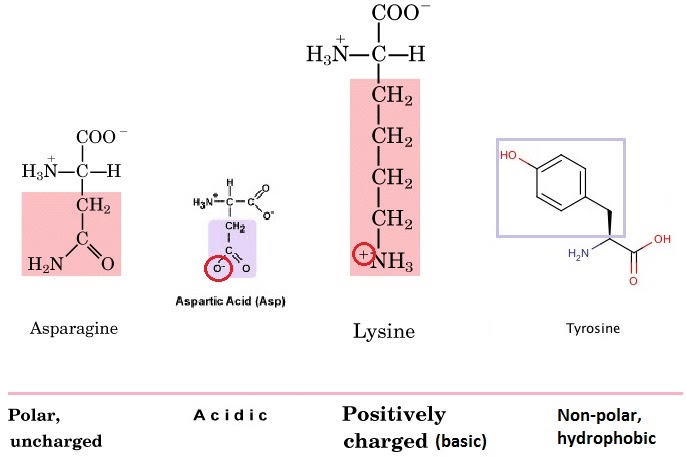
The R group (functional group) on the amino acid also determines the characteristics of that amino acid, including whether it is basic or acidic, polar or hydrophobic. Amino acids can be charged (positively or negatively) or uncharged, polar or non-polar. On asparagine, for example, it is the extra carboxyl (CO) group that makes it polar.
The different extent to which different amino acids are hydrophobic or hydrophilic determines how they behave in the technique called chromatography.