Handling mRNA
Genetic engineering of bacteria
Polymerase chain reaction (PCR)
Visualising DNA with gel electrophoresis
Mining DNA for data
Gene delivery in eukaryotes
Gene technology in research and agriculture
RNA interface (RNAi)
Handling mRNA
As previously covered, mRNA is messenger RNA i.e. the molecule that takes the genetic information encoded by DNA (transcription) and brings it to the ribosome to initiate translation of the code into a polypeptide.
The production of mRNA in human cells is not a simple transcription of DNA, as previously seen. The pre-mRNA is the simple transcript of DNA, but the mature mRNA has to be spliced first, as well as go through a couple of post transcriptional edits before being ready for translation at the ribosome site.
Why is this significant for gene technologies? Gene tech relies on using genetic information in DNA to accomplish various feats. However, eukaryotic DNA contains introns that are not used as code for the desired product. Therefore, a sequence of DNA without introns must be produced.
This is accomplished by using a mature mRNA sequence and decoding it in reverse back to DNA. The enzyme used for this is reverse transcriptase. This produces a usable DNA sequence free of introns (more on this later).
Right then, what are these post transcriptional edits of mRNA?
In eukaryotes, genes contain non-coding sequences which must be removed before mRNA is used to produce proteins. These are called introns as opposed to exons which are coding sequences. Splicing therefore is the process of excising (cutting out) introns to be left with mRNA containing purely coding sequence.
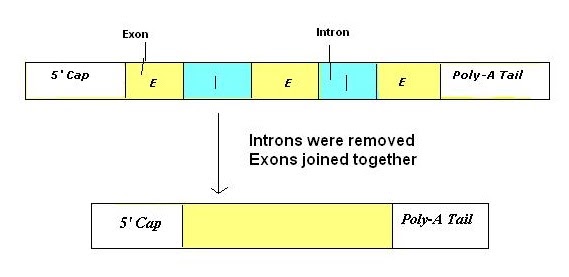
This process can result in several different mRNA products from the same DNA sequence. If the introns and exons are arranged differently, the mRNA will code for different amino acids. It’s termed alternative splicing.
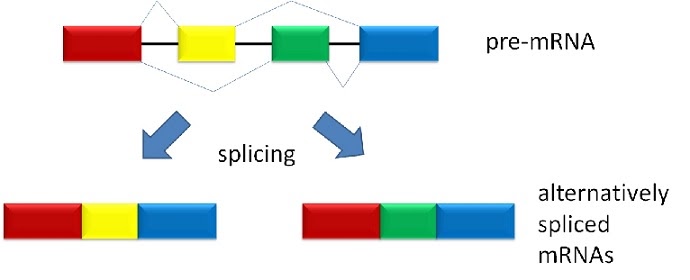
Since these two possible mRNA products code for different amino acids represented by the different colours (red-yellow-blue versus red-green-blue), the resulting protein after translation of mRNA could function differently. If an enzyme, it may affect its ability to catalyse its reactions, or its efficiency. Equally, the change could not make a difference in another scenario at all.
You might have noticed some additional bits to the final mature mRNA in the diagram, called 5′ cap and Poly-A tail. These, alongside spicing…
